The measured mass is converted to a molecule count using information about the fragment length distribution. 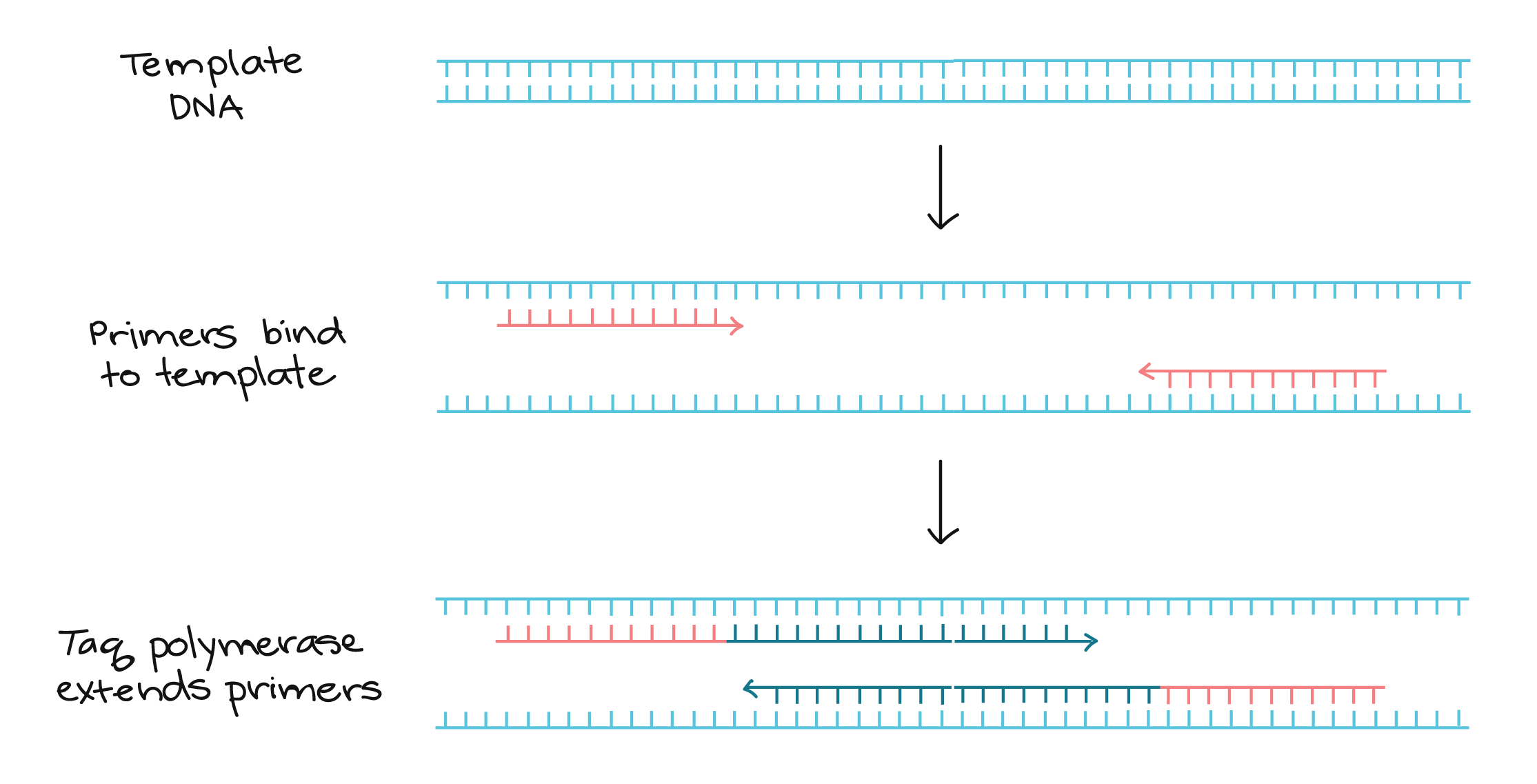
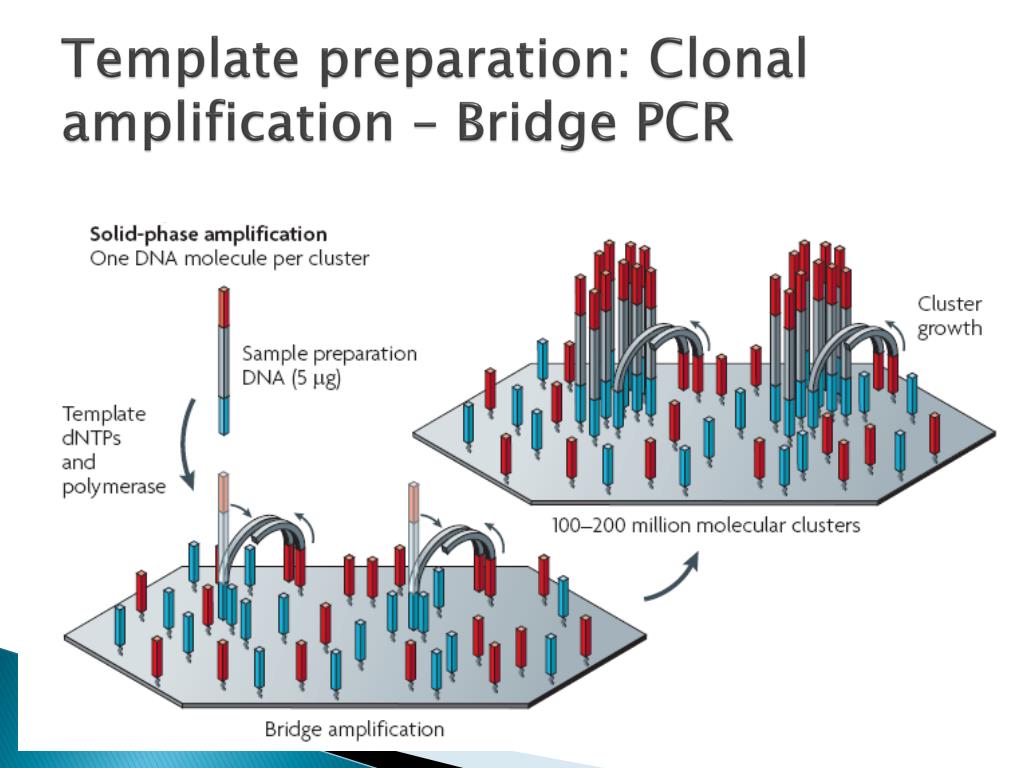
The manufacturers' protocols call for quantification of sequencing libraries by mass using capillary gel electrophoresis or UV spectrophotometry. The accuracy of digital PCR and its ability to count only amplifiable molecules obviate the need for expensive and time-consuming titration sequencing runs. Inaccuracy in quantification is addressed by the manufacturers through 'titration' runs of the sequencer, which are used to empirically divine the concentration of productive DNA fragments in the sequencing library. Accurate quantification of the sequencing library is essential to achieve high yield and high quality sequencing. Overestimation of library concentration results in fewer DNA-bearing beads after emulsion PCR or sparse clustering in bridge PCR, in which case the full capacity of the sequencer cannot be realized. The consequences are mixed signals or un-resolvable clusters, which reduce the number of high quality reads. Underestimation of library concentration results in multiple library molecules associating with the same bead within an emulsion microdroplet or overlapping images of DNA clusters after bridge PCR. On the Illumina/Solexa platform, library molecules are captured by surface-tethered probes complementary to the adaptor sequences and are amplified by bridge PCR to convert a single DNA molecule into a surface-bound cluster with many copies of the same sequence.Īccurate quantification of the number of library molecules is a critical factor affecting next-generation sequencing performance. On the Roche/454 and ABI/SOLiD platforms, emulsion PCR is used to amplify a single DNA molecule to millions of copies of the same sequence all attached to a single polymer bead. The presence of these adaptors enables the amplification of random-sequence inserts by parallel PCR amplification of millions of individual DNA molecules. 
During library creation, adaptor sequences are appended to both ends of the DNA molecules in a sample. The workflow for these new sequencing technologies proceeds as follows: library creation, library quantification, massively parallel clonal PCR amplification of library molecules, and sequencing. A key advance facilitating higher throughput and lower costs for several of these platforms was migration from the clone-based sample preparation used in Sanger sequencing to the massively parallel clonal PCR amplification of sample molecules on beads (Roche 454 and ABI Solid) or on a surface (Solexa). The digital PCR assay allows absolute quantification of sequencing libraries, eliminates uncertainties associated with the construction and application of standard curves to PCR-based quantification, and with a coefficient of variation close to 10%, is sufficiently precise to enable direct sequencing without titration runs.Ī new generation of sequencing technologies based on sequencing by synthesis and sequencing by ligation are revolutionizing biology, biotechnology, and medicine. This study is the first to definitively demonstrate the successful sequencing of picogram quantities of input DNA on the 454 platform, reducing the sample requirement more than 1000-fold without pre-amplification and the associated bias and reduction in library depth. We successfully sequenced low-nanogram scale bacterial and mammalian DNA samples on the 454 FLX and Solexa DNA sequencing platforms. We demonstrate the use of digital PCR to accurately quantify 454 and Solexa sequencing libraries, enabling the preparation of sequencing libraries from nanogram quantities of input material while eliminating costly and time-consuming titration runs of the sequencer. Second, each library requires a titration sequencing run, thereby increasing the cost and lowering the throughput of sequencing. For many applications, including metagenomics and the sequencing of ancient, forensic, and clinical samples, the quantity of input DNA can be critically limiting. 
First, large amounts of sample-typically micrograms-are needed for library preparation, thereby limiting the scope of samples which can be sequenced. This requirement has two unfavorable consequences. Next-generation DNA sequencing on the 454, Solexa, and SOLiD platforms requires absolute calibration of the number of molecules to be sequenced.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
